If the models present in the libraries do not suit your needs, you can write a model yourself. You can either start completely from scratch or adapt a model existing in the libraries. In both cases, the syntax of the Mlxtran language is detailed on the mlxtran webpage. You can also copy-paste models from the mlxtran model example page.
Videos on this page use the application mlxEditor included in previous versions of MonolixSuite. From the 2021R1 version on, the editor is integrated within the interface of Monolix, as shown on the screenshots on this page, and it can also be used as a separate application.
Writing a model from scratch
- 8.case_studies/hcv_project (data = ‘hcv_data.txt’ , model=’hcvNeumann98_model.txt’)
In the Structural model tab, you can click on New model to open the editor integrated within Monolix, and start writing your own model. The new model contains a convenient template defining the main blocks, input parameters and output variables. When you are done, click on Create model button of Monolix to save your new model file. After saving, the model is automatically loaded in the project.
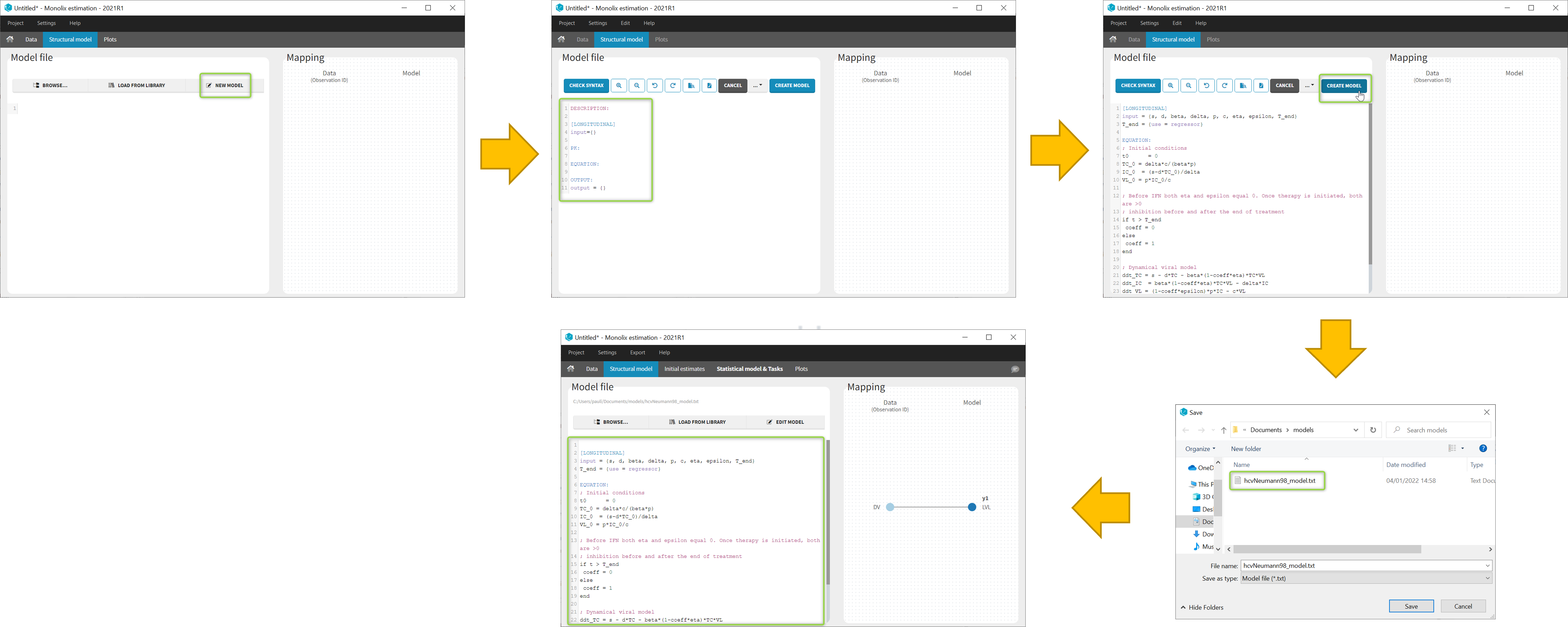
You can even create your own library of models. An example of a basic library which includes several viral kinetics model is available in the demos 8.case_studies/model.
Note: A button “Check syntax” is available to check that there is no syntax error in the model. In case of an error, informative messages are displayed to help correct the error. The syntax check is also automatically applied before saving the model so that only a model with a valid syntax can be saved.
![]()
Understanding the error messages
The error messages generated when syntax errors are present in the model are very informative and quickly help to get the model right. The most common error messages are explained in detail in this video.
Modifying a model from the libraries
Browse existing models from the libraries using the Load from library button, then click the “file” icon next to a model name. This opens a pop-up window where the content of the model file is displayed. Click on Open in editor to open the model file in the MlxEditor. There you can adapt the model, for instance to add a PD model. Be careful to save the new model under a new name, to avoid overwriting the library files.
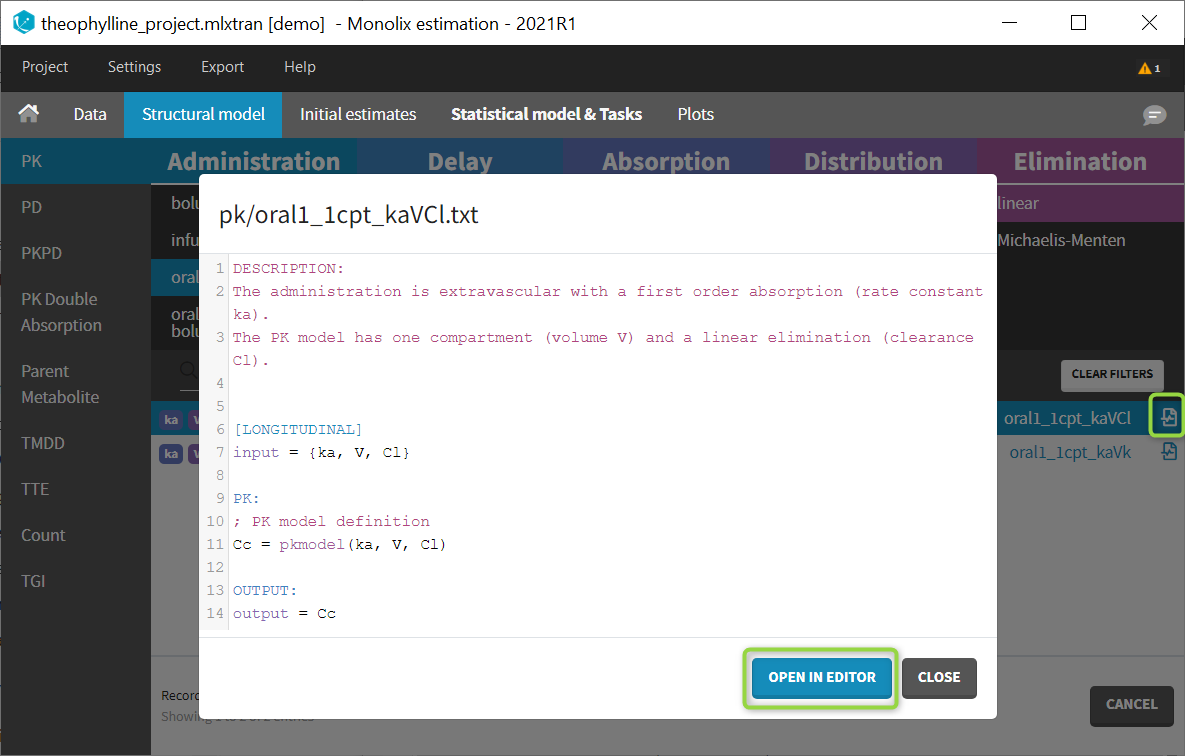
The video below shows an example of how a scale factor can be added:
See the dedicated webpage for more details on model libraries.